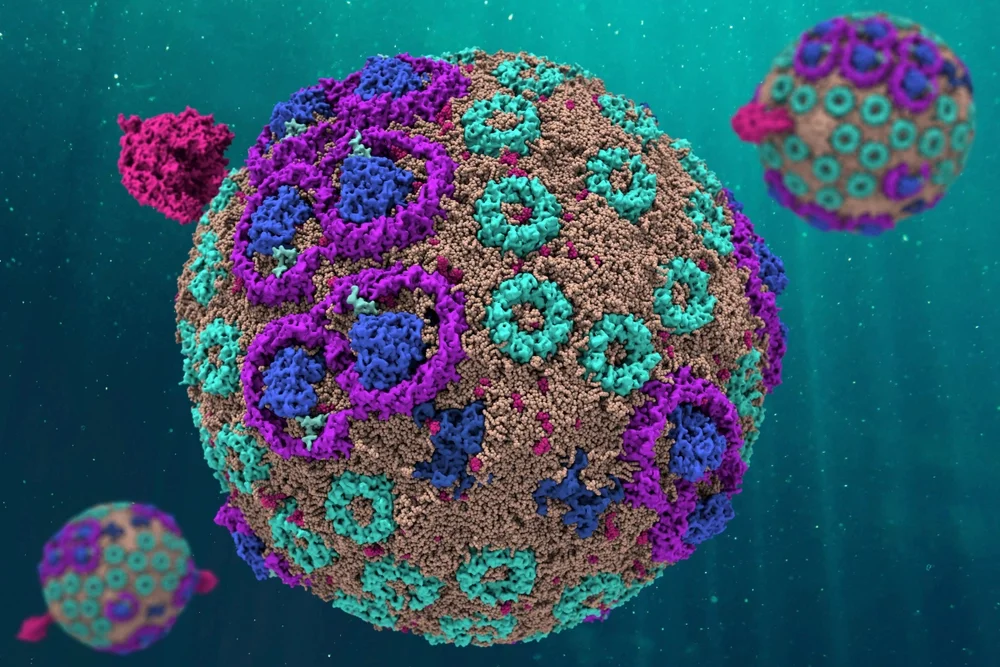
Scientists have simulated every atom of a light-harvesting structure in a photosynthetic bacterium that generates energy for the organism. The simulated organelle behaves just like its counterpart in nature, the researchers report. The work is a major step toward understanding how some biological structures convert sunlight into chemical energy, a biological innovation that is essential to life.
The researchers report their findings in the journal Cell.
The team originally was led by University of Illinois physics professor Klaus Schulten and continued the work after Schulten’s death in 2016. The study fulfills, in part, Schulten’s decades-long dream of discovering the mechanisms by which atomic-level interactions build and animate living systems.
Schulten decided very early in his career to study photosynthetic systems, said study co-author Melih Sener, a research scientist at the U. of I.’s Beckman Institute for Advanced Science and Technology, where much of the work was conducted. Schulten and Sener modeled the chromatophore, a primitive photosynthetic organelle that produces chemical energy in the form of a molecule known as ATP. That work involved a long-term collaboration with Neil Hunter from the University of Sheffield, who provided much of the experimental data.
“Schulten was a physicist; he wanted to understand biology at the physics level,” said Illinois biochemistry professor and study co-author Emad Tajkhorshid. “But then he realized biology only works if you put all of the complexity into the model. And the only way to do that was with supercomputers.”
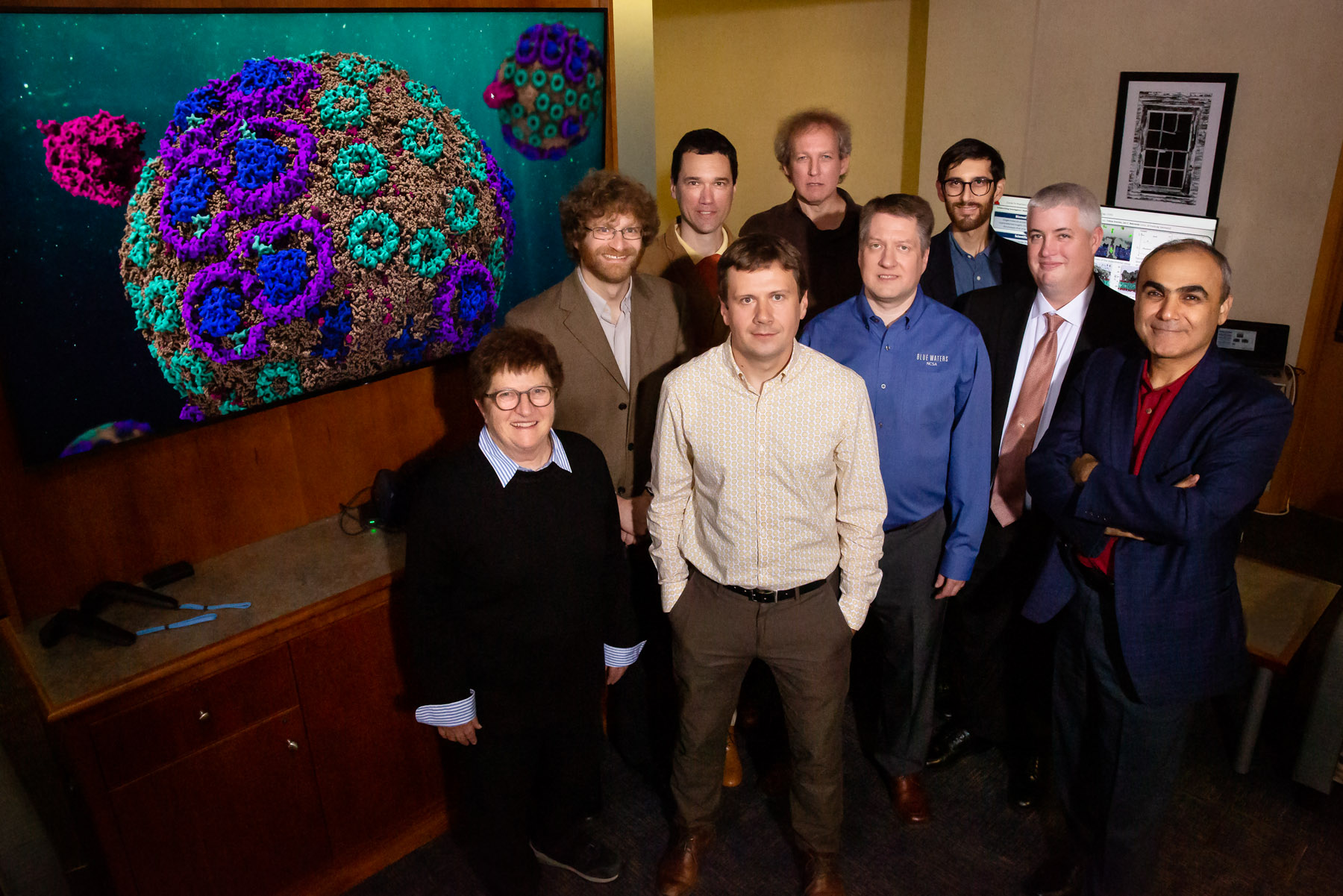

Photo by L. Brian Stauffer
Over the years, Schulten recruited and supported collaborators at Illinois and elsewhere to help him tackle the challenge. The team constructed a 136 million-atom model of the chromatophore, an effort that required a colossal amount of supercomputer power over a period of four years. The work was conducted on the Titan and Summit supercomputers at the Oak Ridge National Laboratory in Knoxville, Tennessee; and on Blue Waters, housed at the National Center for Supercomputing Applications at the U. of I.
Schulten and his colleagues had already conducted molecular simulations of many of the individual protein and lipid components of the chromatophore, which produces the ATP needed to power a living cell.
“The chromatophore has an antenna, a battery and a motor,” said study lead author Abhishek Singharoy. The antenna harvests light, the battery directs that energy to the motor and the motor cranks out ATP, he said. Singharoy worked with Schulten at Illinois before accepting a professorship at Arizona State University, Tempe in 2017.
Figuring out how the system worked required putting all the parts together, said Illinois physics professor Aleksei Aksimentiev, who guided the project to completion after Schulten’s death. This meant dissecting the chromatophore with every tool available to science, from laboratory experiments to electron microscopy, to programming innovations that broke down the computing challenge into manageable steps, Aksimentiev said.
Once they had a working model of the chromatophore, the researchers watched simulations that revealed how the organelle functioned under different scenarios. They changed the concentration of salt in its environment, for example, to see how it coped with stress.
When they exposed their simulated organelle to conditions that it typically experiences in the cell, they were surprised by how it behaved. It immediately became less spherical, and certain proteins embedded in the membrane began to clump together.
“We started with a perfect sphere, but very rapidly it became imperfect, with flat areas and little areas with high curvatures,” Aksimentiev said. “And all of that, our calculations reveal, has a biological role.”
The clumping proteins create patches of positive and negative charges that facilitate the distribution of electrons across the system, the researchers found. The electrons are ultimately swapped for protons, which drive an enzyme known as an ATP synthase, the motor that produces ATP.
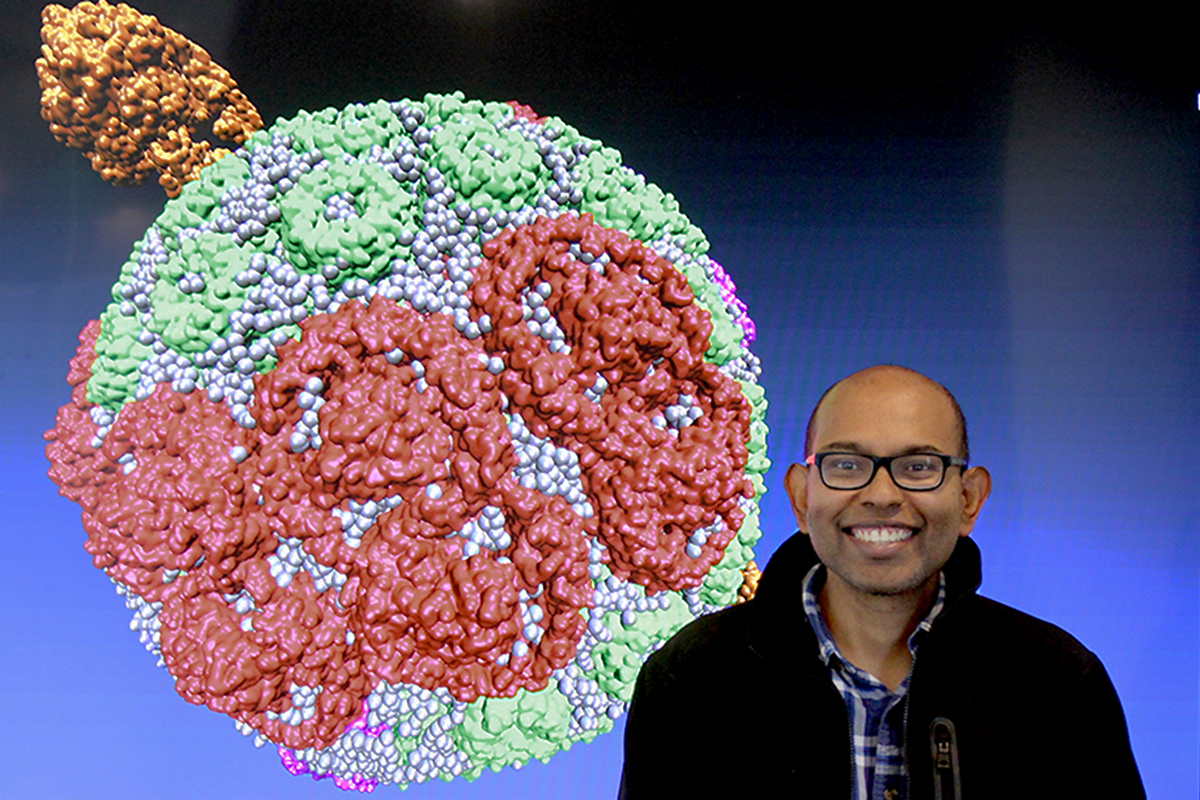
Photo courtesy Abhishek Singharoy
“The chromatophore’s structure is like a circuit diagram,” Sener said. “If you know how the energy and charges travel in it, you know how the machine works. The chromatophore is basically an electronic device.”
The study confirms that, at the atomic scale, physics drives biology, the researchers said. The work will inform future studies of more complex energy-generating organelles in other microorganisms, and in plants and animals, they said. And it will advance scientists’ understanding of nature’s solution to a perpetual human problem: how to efficiently extract energy from the environment without poisoning oneself.
The research was made possible through the National Institutes of Health Center for Macromolecular Modeling and Bioinformatics and the National Science Foundation Center for the Physics of Living Cells, both at the University of Illinois. The Research Corporation for Science Advancement and the Gordon and Betty More Foundation supported the work in Arizona.
Aksimentiev and Tajkhorshid are affiliates of the Beckman Institute at the U. of I.
Diana Yates | Life Sciences Editor, U. of I. News Bureau | 217-333-5802