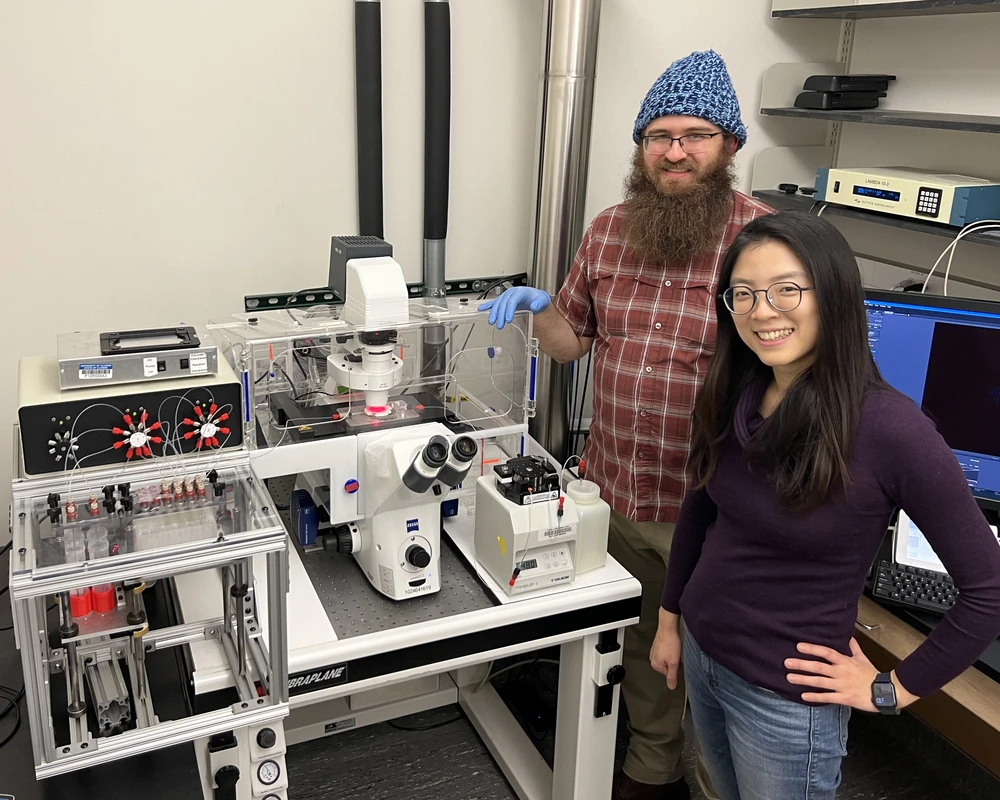

With recent advances in spatial omics technology, scientists are getting closer to realizing an ambitious dream in biology: creating a detailed map of where every single molecule is located in each and every cell within a tissue.
Illinois Professor Hee-Sun Han explains that mapping just the position of cells within a tissue can easily be done, but her research group will be localizing all the molecules within each and every cell within a tissue, which will accomplish two goals:
- Create a comprehensive, detailed spatial map of the molecules within every cell within a tissue.
- Show how each cell, with its own distribution of molecules, is structurally organized in a tissue, which relates to each cell’s function.
Molecular and cellular interactions are mediated by physical contacts; therefore, knowing the position – how molecules and cells are spatially organized within tissue – can further advance our understanding of tissue structure and function, which is a fundamental goal in life sciences research.
Traditional spatial technologies can generate spatial data for only a few molecules at a time, but spatial omics technologies can simultaneously localize hundreds to tens of thousands of molecules, creating a comprehensive spatial map at unprecedented scale and resolution.
Professor Han in the Department of Chemistry is driving technical innovations in spatial omics by further innovating the single molecule resolution spatial transcriptomics platform they established to enable high throughput multimodal imaging. On the analytics front, the Han group is collaborating with biostatisticians and bioinformaticists to develop new analytical methods that extract unique biological insights from such rich spatial datasets.
Recently, Han received more than $2 million in funding from the National Institutes of Health to develop chemical tools to allow sequential imaging of proteins and RNAs at the genome scale in intact tissues.
“The new technology will allow researchers to tease apart the interactions and communication networks at unprecedented details and scale thereby providing critical insights into how cells and molecules collectively perform higher-level functions. For instance, we can much better model how diseased cells perturb nearby cells and spread over time,” Han said.
In addition to driving the technical innovations, Han also has been leading the Spatial Omics Initiative at the Carl R. Woese Institute of Genomic Biology at Illinois along with professor Dave Zhao in the Department of Statistics.
In March 2022, Han, Zhao, and others organized a campus-wide, full day workshop on spatial omics that attracted more than 80 people across the campus from more than 12 departments. Building on that momentum and enthusiasm, the goal of the Spatial Omics Initiative is to make major transformational contributions in this fast-growing field through collaborative, multi-disciplinary efforts that utilize artificial intelligence, machine learning and various science and engineering principles in the fields of biology, chemistry, medicine, and engineering.
“The potential of spatial omics is huge, much bigger than what is shown so far. Having the “Google map” of a cell and a tissue, researchers can build systemic models on how molecular and cellular mechanisms learned from traditional biochemistry are spatially and functionally organized. Furthermore, machine learning and artificial intelligence can extract new, invaluable insights from such rich spatial data. In addition to driving technical advances, I want to serve as a key node in the innovation ecosystem at Illinois,” Han said.
Han is also affiliated with the Center for Biophysics and Quantitative Biology, the Bioengineering and the Neuroscience Program at Illinois.
You can reach Hee-Sun Han at hshan@illinois.edu.