
By Ananya Sen, University of Illinois at Urbana-Champaign
Researchers from the Carl R. Woese Institute for Genomic Biology, including chemistry professor Wilfred van der Donk, collaborated with scientists at Oxford University on a paper recently published in Cell revealing the function of LanCL proteins, which have been found in eukaryotic cells for years but their function was unknown.
The study is the first step towards understanding the importance of these ubiquitous proteins.
Bacteria contain enzymes called LanC that are capable of producing small proteins called lanthipeptides, which are characterized by the addition of a thiol group to a modified serine or threonine amino acid. Similar proteins—called LanC-like or LanCL—have been found in different eukaryotic cells for decades, but their function was unknown.
"LanCLs are found in nearly all higher organisms, including humans. Although scientists have worked on these proteins for over 20 years, we didn't know their function. We had several hypotheses, which we kept ruling out based on our experiments," said Wilfred van der Donk (MMG), Richard E. Heckert Endowed Chair in Chemistry and Howard Hughes Medical Institute Investigator.
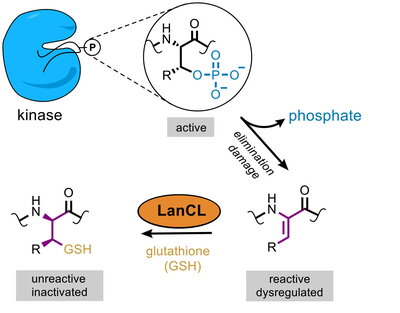
The first breakthrough came in 2015, when the Nair lab in the Department of Biochemistry solved the crystal structure of a LanC-containing protein in bacteria. The protein was bound to another enzyme called a kinase, which modifies proteins by adding a phosphate group. Inspired by this discovery, the researchers tested whether LanCL proteins were also binding to kinases in eukaryotic cells. "We saw that they were able to bind to many kinases, including AKT and mTOR, and all of a sudden the pieces of the puzzle started forming a picture," van der Donk said.
The next piece fell into place in collaboration with Benjamin Davis, a professor of chemistry at the University of Oxford. The Davis group showed that eliminating a particular phosphate group in kinases causes them to become activated. Scientists had assumed that such processed proteins would be inactive. Together, the Illinois and Oxford groups were able to show that LanCL adds glutathione to kinases with eliminated phosphate groups, after which the kinases became deactivated. "We realized that when the LanCL proteins are absent, the cell has a big problem because there are active proteins floating around that need to be turned off," van der Donk said.
The importance of these proteins became evident in mice that lacked them. "A third of the mice that lack these enzymes die when they are between four to six months old. They die suddenly without getting sick and we still don't understand why," said Jie Chen (GNDP), a professor of cell and developmental biology.
The researchers are interested in understanding the role of these proteins and making a complete list of all the possible targets of LanCLs. "When you have abnormal kinases, it can cause all kinds of problems, including cancer. LanCL proteins eliminate these damaged kinases and it is possible that they also affect other proteins that we are not aware of. We need to connect their cellular functions to the results we saw in the mice," Chen said.
"This study is just the tip of the iceberg. Since these proteins are found everywhere, you can also imagine their effects in feedstock and the future of farming," said Satish Nair (MME/MMG), Head of the Department of Biochemistry.
"This study was possible because of the persistence of our graduate students. Most of us would have given up long ago because the studies were initially going nowhere," Nair said. "It also shows the importance of exploratory research, where you're essentially just looking around. Although it is risky, it is great to see that there are rewards for students who stick it out," van der Donk said.
The paper "LanCLs add glutathione to dehydroamino acids generated at phosphorylated sites in the proteome" was published in Cell.
More information: Kuan-Yu Lai et al, LanCLs add glutathione to dehydroamino acids generated at phosphorylated sites in the proteome, Cell (2021). DOI: 10.1016/j.cell.2021.04.001
Read more about this research: Piecing together a puzzle in protein damage - Rosalind Franklin Institute (rfi.ac.uk)